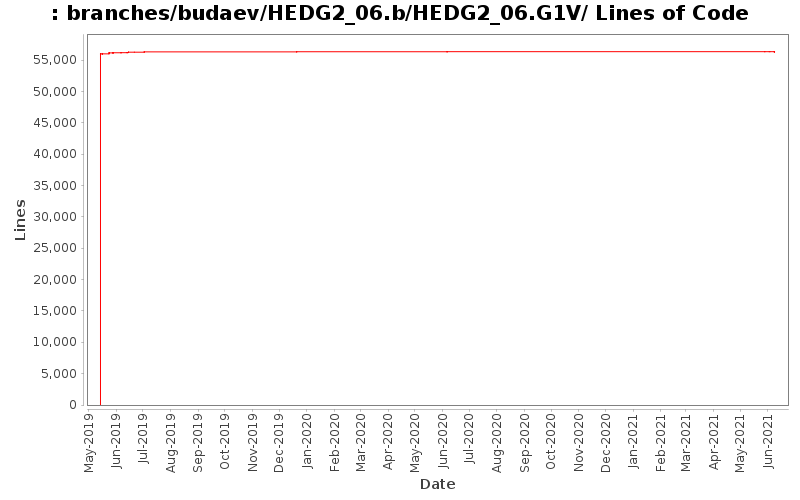
[root]/branches/budaev/HEDG2_06.b/HEDG2_06.G1V
![]() dox
(126 files, 319170 lines)
dox
(126 files, 319170 lines)
![]() pfunit
(2 files, 77 lines)
pfunit
(2 files, 77 lines)
![]() tests
(6 files, 1831 lines)
tests
(6 files, 1831 lines)
![]() tools
(2 files, 135 lines)
tools
(2 files, 135 lines)
| Author | Changes | Lines of Code | Lines per Change |
|---|---|---|---|
| sbu062 | 51 (100.0%) | 56578 (100.0%) | 1109.3 |
`O2` and higher optimization does not work on Windows Intel Fortran, propagate fix r11234
6 lines of code changed in 1 file:
delete dependence of living cost on smr, very small effect and potentially slow down simulation
Note: this applies changes r11231:r11232
applied diff `svn diff -r 11226:11232 m_body.f90` for all branches
2 lines of code changed in 1 file:
fix exponent as in r11226
2 lines of code changed in 1 file:
repeat r11209: need even longer record to fit all values
3 lines of code changed in 1 file:
need extra length for the record to avoid writing data to non-existent space beyond last character, useful against huge history crashing
1 lines of code changed in 1 file:
apply patch from simulation, worked well, Limit maximum level of sex hormones to 10.0 to avoid huge growth of it and reproductive beyond limits
Patch here:
{{{
Index: m_body.f90
===================================================================
--- m_body.f90 (revision 10934)
+++ m_body.f90 (working copy)
@@ -1439,15 +1439,15 @@
if (length_increment > TOLERANCE_LOW_DEF_SRP) then
if (this%is_male()) then
!> - If the agent is **male**, **testosterone** is incremented.
- call this%testosterone_set( &
+ call this%testosterone_set( min( 10.0_SRP, &
this%testosterone_get() + &
- this%testosterone_get() * steroid_increment_factor )
+ this%testosterone_get() * steroid_increment_factor ) )
else
!> - If the agent is **female**, **estrogen** is incremented.
!! .
- call this%estrogen_set( &
+ call this%estrogen_set( min( 10.0_SRP, &
this%estrogen_get() + &
- this%estrogen_get() * steroid_increment_factor )
+ this%estrogen_get() * steroid_increment_factor ) )
end if
!> If there was no growth and the gonadal steroids are not incremented,
!! the current values are still saved in the history stack by calling
}}}
4 lines of code changed in 1 file:
fix power factor
2 lines of code changed in 1 file:
white space only: fix if block indent for readability
1 lines of code changed in 1 file:
gamma2gene: merged tested r9487: merged changes in (allelescale) and (gamma2gene) backends from SIGMOID_tweak_new r9486
*Patch code:*
{{{
Index: m_common.f90
===================================================================
--- m_common.f90 (.../HEDG2_06.b2/HEDG2_06.P1H) (revision 9189)
+++ m_common.f90 (.../HEDG2_06.b3/HEDG2_06.P1H) (revision 9550)
@@ -2594,6 +2594,11 @@
!! `commondata::allelescale()` functions.
integer, parameter, public :: ALLELERANGE_MAX = 10000
+ !> Conversion parameter that defines the scaling of the integer allele
+ !! values ::ALLELERANGE_MIN to ALLELERANGE_MAX are converted to ::zero to
+ !! this parameter value as the maximum. See ::allelescale() for details.
+ real(SRP), parameter, public :: ALLELESCALE_MAX = 20.0_SRP
+
!> Number of additive allele components
integer, parameter, public :: ADDITIVE_COMPS = 3
@@ -6875,14 +6880,15 @@
!! @param[in] raw_value raw input value, integer within
!! `commondata::allelerange_min` and `commondata::allelerange_max`
!! @retval Returns the value of conversion function: integer alleles
- !! to real internal value 0 to 1
+ !! to real internal value ::zero to ::allelescale_max
!! @details Allele conversion function for the relationship between the
!! genome integer allele value and its converted real value in
!! the neuronal response function. The function rescales integer
!! allele value @f$ I_{i} @f$ within the range
!! @f$ [I_{min},I_{max}] @f$ to real values @f$ r @f$ within the
- !! range @f$ [0,1] @f$ using the formula:
- !! @f[ r = \frac{I - I_{min}}{I_{max}-I_{min}} @f]
+ !! range @f$ [0,M] @f$, where @f$ M @f$ is defined by the
+ !! ::allelescale_max parameter. Conversion is performed by the
+ !! ::rescale() backend function.
!! See implementation notes on `allele_value` component of the
!! `GENE` derived type.
!! @note Note that it is an *elemental function* that can accept both
@@ -6889,20 +6895,13 @@
!! scalar and vector arguments.
elemental function allelescale(raw_value) result(converted)
- ! @param[in] raw_value raw input value, integer within
- ! `commondata::allelerange_min` and `commondata::allelerange_max`.
integer, intent(in) :: raw_value
- ! @retval Returns the value of conversion function: integer alleles
- ! to real internal value 0 to 1
real(SRP) :: converted
- converted = (real(raw_value,SRP) - ALLELERANGE_MIN) / &
- (ALLELERANGE_MAX - ALLELERANGE_MIN)
+ converted = rescale( real(raw_value,SRP), &
+ real(ALLELERANGE_MIN,SRP), real(ALLELERANGE_MAX,SRP),&
+ ZERO, ALLELESCALE_MAX )
- !> @warning We must never accept zero values of alleles as they will
- !! result in **division by zero** in `commondata::gamma2gene()`.
- if (converted < ZERO) converted = ZERO
-
end function allelescale
!-----------------------------------------------------------------------------
@@ -7025,13 +7024,13 @@
! function.
! @note Note that the raw integer gene values are accepted by this
! function as `allelescale` is called automatically inside.
- integer, dimension(ADDITIVE_COMPS), intent(in) :: gs
+ integer, dimension(:), intent(in) :: gs
! @param[in] gh half-max effect: Gene/constant for the signal strength
! giving half max effect.
! @note Note that the raw integer gene values are accepted by this
! function as allelescale is called automatically inside.
- integer, dimension(ADDITIVE_COMPS), intent(in) :: gh
+ integer, dimension(:), intent(in) :: gh
! @param[in] signal perception: Input value of (external or internal)
! stimulus perception.
@@ -7045,7 +7044,7 @@
! Local variables:
real(SRP) :: perception
- real(SRP) :: d1
+ real(HRP) :: d1, nr
integer :: i
! Parameter that specifies the minimum perception. Can be commondata::zero
@@ -7065,7 +7064,7 @@
if (present(erpcv)) then
if (erpcv > TOLERANCE_HIGH_DEF_SRP) then
perception = max(FORCED_MIN_PERCEPT, RNORM(signal,(erpcv*signal)**2))
- else
+ else
perception = max(FORCED_MIN_PERCEPT, signal)
end if
else
@@ -7072,16 +7071,22 @@
perception = max(FORCED_MIN_PERCEPT, signal)
end if
- neuronal_response = 0.0_SRP
+ nr = 0.0_HRP
! @warning The `do concurrent` construct is F2008 and can not (yet) be
! implemented in all compilers. Use normal `do` in such a case.
- do concurrent (i=1:ADDITIVE_COMPS)
- d1 = ( perception / alleleconv( allelescale(gh(i)) ) &
- ) ** alleleconv( allelescale(gs(i)) )
- neuronal_response = neuronal_response + d1/(1.0_SRP+d1)
+ do concurrent (i=1:size(gs))
+ d1 = (real(perception, HRP) / real(alleleconv(allelescale(gh(i))),HRP)) &
+ ** real( alleleconv( allelescale(gs(i)) ), HRP )
+ nr = nr + d1/(1.0_HRP+d1)
end do
+ if (nr < ZERO) then
+ neuronal_response = ZERO
+ else
+ neuronal_response = real(nr, SRP)
+ end if
+
end function gamma2gene_additive_i4
!-----------------------------------------------------------------------------
@@ -7130,13 +7135,13 @@
! function.
! @note Note that the raw integer gene values are accepted by this
! function as `allelescale` is called automatically inside.
- real(SRP), dimension(ADDITIVE_COMPS), intent(in) :: gs
+ real(SRP), dimension(:), intent(in) :: gs
! @param[in] half-max effect: Gene/constant for the signal strength giving
! half max effect.
! @note Note that the raw integer gene values are accepted by this
! function as `allelescale` is called automatically inside.
- real(SRP), dimension(ADDITIVE_COMPS), intent(in) :: gh
+ real(SRP), dimension(:), intent(in) :: gh
! @param[in] perception: Input value of (external or internal) stimulus
! perception.
@@ -7150,7 +7155,7 @@
! Local variables:
real(SRP) :: perception
- real(SRP) :: d1
+ real(HRP) :: d1, nr
integer :: i
! Parameter that specifies the minimum perception. Can be commondata::zero
@@ -7169,7 +7174,7 @@
if (present(erpcv)) then
if (erpcv > TOLERANCE_HIGH_DEF_SRP) then
perception = max(FORCED_MIN_PERCEPT, RNORM(signal,(erpcv*signal)**2))
- else
+ else
perception = max(FORCED_MIN_PERCEPT, signal)
end if
else
@@ -7176,16 +7181,22 @@
perception = max(FORCED_MIN_PERCEPT, signal)
end if
- neuronal_response = 0.0_SRP
+ nr = 0.0_HRP
! @warning The `do concurrent` construct is F2008 and can not (yet) be
! implemented in all compilers. Use normal `do` in such a case.
- do concurrent (i=1:ADDITIVE_COMPS)
- d1 = (perception / alleleconv(gh(i)) &
- ) ** alleleconv(gs(i))
- neuronal_response = neuronal_response + d1/(1.0_SRP+d1)
+ do concurrent (i=1:size(gs))
+ d1 = (real(perception,HRP) / real(alleleconv(gh(i)),HRP)) &
+ ** real( alleleconv(gs(i)),HRP )
+ nr = nr + d1/(1.0_HRP+d1)
end do
+ if (nr < ZERO) then
+ neuronal_response = ZERO
+ else
+ neuronal_response = real(nr, SRP)
+ end if
+
end function gamma2gene_additive_r4
!-----------------------------------------------------------------------------
}}}
42 lines of code changed in 1 file:
portability, merged r9188, as in HEDTOOLS r9185
71 lines of code changed in 1 file:
merged r8486, save all agent data at each time step in (lifecycle_preevol)
- tests passed
52 lines of code changed in 2 files:
foxed global bug causing GA_REPRODUCE_N_MIN reproducing sub-population in all cases
2 lines of code changed in 2 files:
(LOG_IEEE): log IEEE signalling, backport from r8442:r8443
54 lines of code changed in 1 file:
shorter log message
1 lines of code changed in 1 file:
backported r8427:r8428 IPO now works in ifort
3 lines of code changed in 1 file:
backported (ieee_error_reporting) r8421:r8423
41 lines of code changed in 2 files:
backported reverse sorfing in (qsort) as in r8409
13 lines of code changed in 1 file:
backported fix r8348
1 lines of code changed in 1 file:
backported r8339:r8344 from parent model
- r8344 optional argument for checking only invalid values
- r8343 checks for Inf
30 lines of code changed in 1 file:
backported r8339 IEEE_IS_NAN() is siandard
3 lines of code changed in 1 file:
fix doxygen comments after backporting
- forgotten on parameters descriptions
4 lines of code changed in 1 file:
more conformant explanations for (do_sanitise) and (ieee_error_reporting)
- more correct doxygen style
45 lines of code changed in 1 file:
backported latest changes IEEE exceptions from HEDG2_06 r8320
183 lines of code changed in 2 files:
**compiler bug**, debug: fix for intel **ifort 19** compiler bug
- accessor function `is_available()` is not accessible
resulting in crashing in high cpu-optimised build
(`-O3` etc)
5 lines of code changed in 1 file:
debug: more information logging
- log value `distance` and `visrange` in `LOG_DBG` because
they occur quite often, need checking actual values if
they are really not very much different, small
differences can be ignored in log WARNINGs
2 lines of code changed in 1 file:
popsize increased to 5000
1 lines of code changed in 1 file:
merged fix r8280 from base code **HEDG02_06** to branches
- `make tests` passed
17 lines of code changed in 1 file:
run: all HEDG2_06 model run types placed in sepaerate branches under `HEDG2_06.b`
- `HEDTOOLS` also copied to allow stand-alone unchanged building
55987 lines of code changed in 20 files: